Noncoding RNAs and Cancer
-
Moslemi Naeini, Mozhgan
-
Reproductive Biotechnology Research Center, Avicenna Research Institute, ACECR , Tehran, Iran
-
 M. Ardekani, Ali
Ph.D., Reproductive Bio-technology Research Center, Avicenna Research Institute, ACECR, Tehran, Iran, P.O. Box: 19615-1177, Tel: +98 21 22432020, Fax: +98 21 22432021, E-mail: Ardekani@avicenna.ac.ir
M. Ardekani, Ali
Ph.D., Reproductive Bio-technology Research Center, Avicenna Research Institute, ACECR, Tehran, Iran, P.O. Box: 19615-1177, Tel: +98 21 22432020, Fax: +98 21 22432021, E-mail: Ardekani@avicenna.ac.ir
-
Reproductive Biotechnology Research Center, Avicenna Research Institute, ACECR , Tehran, Iran
Abstract: The eukaryotic complexity involves the expression and regulation of genes via RNA-DNA, RNA-RNA, DNA-protein and RNA-protein interactions. Recently, the role of RNA molecules in the regulation of genes in higher organisms has become more evident, especially with the discovery that about 97% of the transcriptional output in higher organisms are represented as noncoding RNAs: rRNA, snoRNAs, tRNA, transposable elements, 5' and 3' untranslated regions, introns, intergenic regions and microRNAs. MicroRNAs function by negatively regulating gene expression via degradation or translational inhibition of their target mRNAs and thus participate in a wide variety of physiological and pathological cellular processes including: development, cell proliferation, differentiation, and apoptosis pathways. MicroRNA expression profiles in many types of cancers have been identified. Recent reports have revealed that the expression profiles of microRNAs change in various human cancers and appear to function as oncogenes or tumor suppressors. Abnormal microRNA expression has increasingly become a common feature of human cancers. In this review, we summarize the latest progress on the involvement of microRNAs in different types of cancer and their potential use as potential diagnostic and prognostic tumor biomarkers in the future.
Introduction :
Ambros et al found lin-4, a gene which controls the timing of larval development of Caenorhabditis elegans (C. elegans) in 1993. But, the product of this gene was not any protein; instead it produced a pair of small RNAs (1). The longer RNA is 70 nucleotides (nt) that can shape a stem-loop structure and is the precursor of the shorter RNA (22 nt) that is now known as a member of the class of microRNA genes (2). MicroRNAs (miRNAs) are a novel class of endogenous small, non-coding RNAs (ncRNAs) that control gene expression by targeting specific mRNAs for degradation and/or translational repression (3). At the time it appeared that lin-4 was restric-ted to C. elegans because of lack of homology with other species. However, when the second miRNA gene (let-7) with its target miRNA gene (lin-41), was discovered in 2000, it became clear that these miRNA genes are conserved in many species (4). After that, a great number of microRNAs have been identified in mammals (5).
Many members of ncRNAs are part of the functional classes such as miRNAs and small nucleolar RNAs (snoRNAs) that are well conserved in various species. This is because: a) RNAs in both classes function by hybridi-zation to other nucleic acids b) many of the miRNAs have different cellular targets (6) that confine their chance for sequence co-variation and tendency to evolution. Regarding snoRNAs, they encode sequences and struc-tures that mediate binding of compatible RNA-modifying enzymes to the snoRNA tar-get RNA complex, and this further limits se-quence change (7).
miRNAs along with a large set of ncRNAs are known as ‘‘gene regulators’’ that include: Air, H19, Ipw, NTT, Tsix and XIST in mammals and have various functions from potential involvement in the imprinting pro-cess to X-chromosome inactivation in mam-mals (8). Multiple steps and specific cellular machinery are involved in the biogenesis of miRNAs (9). The miRNAs are encoded as short inverted repeats having a double-stranded RNA (dsRNA) stem loop about 70 bp long and are found in both introns and intergenic clusters in the genome (9). The introns and exons of both protein-coding and non coding transcripts are synthesized by RNA polymerase II from where miRNAs are derived (10). In the nucleus, miRNAs are transcribed as primary pri miRNA transcripts, and then they are processed to shape the precursor pre-miRNA stem loop structure and then is transported to the cytoplasm by RanGTP and Exportin 5 where they are cleaved by the Dicer RNAase III endonuclease and produce mature miRNA (21-23 nt) (11).
Human genome and microRNAs
Development, differentiation, growth, and metabolism are regulated by miRNAs and about 500 known miRNA genes are reported to be encoded by the mammalian genome (12). It is estimated that miRNAs regulate about one third of the genes in human genome (13). How the transcription of miRNAs are regula-ted in the cell is not precisely known However, the transcription of miRNAs are known to depend on their localization within the genome and their proximity to host genes and their locations in introns of coding genes, noncoding genes and exons (14). Recent studies have shown that miRNAs are organized in clusters and share the same transcriptional regulation and if having their own promoters, miRNAs are independently expressed (15, 16).
Approximately 50% of miRNAs are trans-cribed from non-protein-coding genes, and the rest are in the introns of coding genes (12). In higher organisms, about 97% of the trans-criptional product is non-coding RNA (ncRNA) which consist of rRNA, tRNA, introns, 5’ and 3’ untranslated regions, trans-posable elements, and intergenic regions, and a large family known as microRNAs, some of which can down-regulate large numbers of target mRNAs (3, 17). Recently, a number of mammalian miRNAs have been reported to be derived from DNA repeats and transposons (18). This finding has lead to a re-evaluation of the functional role of transposons, esp
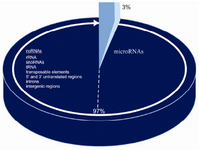
Figure 1. Coding and Non-coding DNA in human genome
|
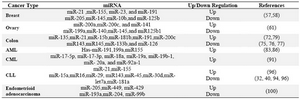
Table 1. Cancer-related miRNAs
|

Table 2. Cancer-related miRNAs
|
|