The Role of MicroRNAs in Human Diseases
-
 M. Ardekani, Ali
Ph.D., Reproductive Biotechnology Research Center, Avicenna Research Institute, ACECR, Tehran, Iran, P.O. Box: 19615-1177., Tel: +98 21 22432020 Fax: +98 21 22432021 E-mail:Ardekani@avicenna.ac.ir
M. Ardekani, Ali
Ph.D., Reproductive Biotechnology Research Center, Avicenna Research Institute, ACECR, Tehran, Iran, P.O. Box: 19615-1177., Tel: +98 21 22432020 Fax: +98 21 22432021 E-mail:Ardekani@avicenna.ac.ir
-
Reproductive Biotechnology Research Center, Avicenna Research Institute, ACECR , Tehran, Iran
Abstract: MicroRNAs (miRNAs) are short RNA molecules which bind to target mRNAs, resulting in translational repression and gene silencing and are found in all eukaryotic cells. Approximately 2200 miRNA genes have been reported to exist in the mammalian genome, from which over 1000 belong to the human genome. Many major cellular functions such as development, differentiation, growth, and metabolism are known to be regulated by miRNAs. Proximity to other genes in the genome and their locations in introns of coding genes, noncoding genes and exons have been reported to have a major influence on the level of gene expressions in eukaryotic cells. miRNAs are well conserved in eukaryotic system and are believed to be an essential and evolutionary ancient component of gene regulatory networks. Therefore, in recent years miRNAs have been studied as a likely candidate for involvement in most biologic processes and have been implicated in many human diseases.
Background :
Since the first draft of human genome was published in February 2001, many new dis-coveries have been made which have eluci-dated the complexity of the human genome and subsequently the human proteome. In the past decade, application of genomics and pro-teomics technologies for early detection of diseases have demonstrated that many types of diseases can be diagnosed at an early stage which would be helpful in initiation of treat-ment protocols at an earlier time point in the clinic. We have previously reported the appli-cation of proteomics technologies for early detection of ovarian cancer (1, 2, 3), prostate cancer (4), lymphatic vascular system (5) and drug-induced cardiac toxicities (6). Other genomics and proteomics technologies have been employed in a variety of other diseases (7, 8) in the past decade.
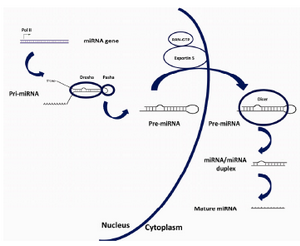
Figure 1. The biosynthesis pathway for miRNAs
|
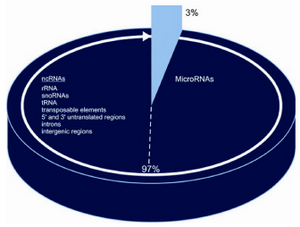
Figure 2. Coding and non-coding DNA in human genome
|
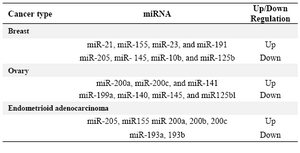
Table 1. miRNAs in reproductive cancers (9)
|
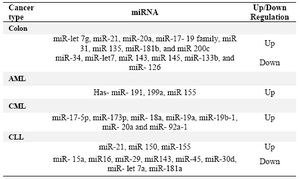
Table 2. miRNAs in cancer (9)
|
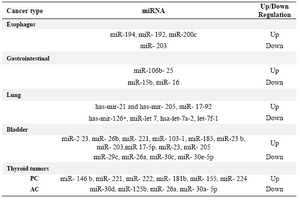
Table 3. miRNAs in cancer (9)
|
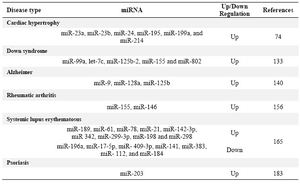
Table 4. miRNAs in human diseases
|
|