Effect of Removal of Spermatogonial Stem Cells (SSCs) from In Vitro Culture on Gene Expression of Niche Factors in Bovine
-
Akbarinejad, Vahid
-
Department of Theriogenology, Faculty of Veterinary Medicine, University of Tehran, Tehran, Iran
-
Theriogenology Association, Faculty of Veterinary Medicine, University of Tehran, Tehran, Iran
-
 Tajik, Parviz
Faculty of Veterinary Medicine, University of Tehran, Tehran, Iran, Tel/Fax: +98 21 61117001 E-mail: ptajik@ut.ac.ir
Tajik, Parviz
Faculty of Veterinary Medicine, University of Tehran, Tehran, Iran, Tel/Fax: +98 21 61117001 E-mail: ptajik@ut.ac.ir
-
Theriogenology Association, Faculty of Veterinary Medicine, University of Tehran, Tehran, Iran
-
Movahedian, Mansooreh
-
Department of Anatomical Sciences, Faculty of Medical Sciences, Tarbiat Modares University, Tehran, Iran
-
Youssefi, Reza
-
Department of Theriogenology, Faculty of Veterinary Medicine, University of Tehran, Tehran, Iran
-
Theriogenology Association, Faculty of Veterinary Medicine, University of Tehran, Tehran, Iran
Abstract: Background: Niche cells, regulating Spermatogonial Stem Cells (SSCs) fate are believed to have a reciprocal communication with SSCs. The present study was conducted to evaluate the effect of SSC elimination on the gene expression of Glial cell line-Derived Neurotrophic Factor (GDNF), Fibroblast Growth Factor 2 (FGF2) and Kit Ligand (KITLG), which are the main growth factors regulating SSCs development and secreted by niche cells, primarily Sertoli cells.
Methods: Following isolation, bovine testicular cells were cultured for 12 days on extracellular matrix-coated plates. In the germ cell-removed group, the SSCs were removed from the in vitro culture using differential plating; however, in the control group, no intervention in the culture was performed. Colony formation of SSCs was evaluated using an inverted microscope. The gene expression of growth factors and spermatogonia markers were assessed using quantitative real time PCR.
Results: SSCs colonies were developed in the control group but they were rarely observed in the germ cell-removed group; moreover, the expression of spermatogonia markers was detected in the control group while it was not observed in the germ cell-removed group, substantiating the success of SSCs removal. The expression of Gdnf and Fgf2 was greater in the germ cell-removed than control group (p<0.05), whereas the expression of Kitlg was lower in the germ cell-removed than control group (p<0.05).
Conclusion: In conclusion, the results revealed that niche cells respond to SSCs removal by upregulation of GDNF and FGF2, and downregulation of KITLG in order to stimulate self-renewal and arrest differentiation.
Introduction :
Spermatogenesis is a complex and tightly-regulated process of cell proliferation and differentiation leading to production of mature spermatozoa from Spermatogonial Stem Cells (SSCs) 1. The coordination of SSCs self-renewal and differentiation is regulated by growth factors produced by somatic cells surrounding SSCs, particularly Sertoli cells 1. In this context, Glial cell line-Derived Neurotrophic Factor (GDNF), Fibroblast Growth Factor 2 (FGF2) and Kit Ligand (KITLG) are of the most important growth factors which have been identified to play pivotal role in regulation of SSCs development 2-4. GDNF, a member of transforming growth factor β family, has been shown to contribute to SSCs self-renewal in different species of mammals including bovine 2,5-9. FGF2 has also been indicated to stimulate germ cells proliferation and self-renewal 3,7. Conversely, KITLG has been found to play a pivotal role in SSCs differentiation 4,10.
Extracellular Matrix (ECM) consisting of a complex network of macromolecules with structural and functional properties is the other part of niche 11,12. The culture of cells on ECM provides an environment partly resembling the normal tissue, which allows to better study the mechanisms underlying the regulation of SSCs in vitro 13-16.
The communication between stem cells and their niche is believed to be reciprocal, meaning that stem cells could also influence the function of niche, culminating in tissue homeostasis 17. In this regard, it has been indicated that following testicular damage and the resultant germ cell elimination, self-renewal is favored over differentiation during the course of regeneration of the germ cell population of seminiferous tubules in mice 18. Although mechanisms underlying this tendency toward self-renewal are not completely understood, overexpression of GDNF 19-21 and insignificant decrease in the Kitlg mRNA expression 22 have been observed to follow elimination of germ cells from seminiferous tubules in mice. To our knowledge, however, whether the absence of SSCs could impact the expression of factors regulating SSCs development is not well-known in bovine. Indeed, studies assessing the changes in molecular behavior of niche cells following elimination of SSCs could help us expand our knowledge in terms of mechanisms whereby niche cells endeavor to repopulate the germ cells following testicular damage, which could further help develop therapies to replenish testicular germ cell in case of testicular injury and the subsequent germ cell depletion 23. Furthermore, this knowledge could guide us to provide an optimal environment-in terms of growth factors-for in vitro culture of SSCs to augment self-renewal since one of the cornerstones for enhancing the efficiency of SSCs transplantation is the enrichment of the donor-derived testicular cell suspension prior to injection into the recipient seminiferous tubules due to the rarity of SSCs in mammalian testes 24.
Accordingly, in the present research, the effect of SSCs removal from in vitro culture on the expression of Gdnf, Fgf2 and Kitlg, as the main niche factors contributing to regulation of SSCs function 2-4 was assessed. ECM was chosen for in vitro culture to provide an environment partly akin to natural condition for testicular cells.
Materials and Methods :
Animals and testicular biopsy: Animal Ethics Committee at University of Tehran approved the present study in terms of animal welfare and ethics. Testicular biopsies were obtained from Holstein calves (aged 3 to 5 months; n=3) as previously described 17. Following testicular biopsy, the specimen was transferred on ice to the laboratory within 2 hr.
Cell isolation: Cell isolation was implemented using a two-step enzymatic isolation procedure 25. The testicular tissue was washed three times in DMEM containing antibiotics and was minced into small pieces by a sterile scissor. The minced testicular tissue was incubated in DMEM containing 1 mg/ml collagenase (Sigma-Aldrich, USA), 1 mg/ml hyaluronidase (Sigma-Aldrich, USA), 1 mg/ml trypsin (Sigma-Aldrich, USA) and 5 µg/ml DNase (Fermentas, Germany) at 37°C in a shaker incubator with 80 cycles per minute for approximately 60 min. The digested testicular tissue was washed three times with DMEM and the supernatant was disposed after each washing, leading to isolation of seminiferous tubules. During the second step of enzymatic digestion, the seminiferous tubules were incubated at 37°C in DMEM containing 1 mg/ml collagenase, 1 mg/ml hyaluronidase and 5 µg/ml DNase until disintegration of the seminiferous tubules and separation of the constituent cells. Individual cells were isolated from the remaining tubule fragments by centrifugation at 30 g for 2 min. Following filtration through 77 and 55 mm nylon filters, the cells were pelleted. The pellet was re-suspended in the DMEM containing antibiotics and 10% knock-out serum replacement (KSR, Gibco, USA).
Cell culture: Six-well plates were coated with extracellular matrix (ECM) gel (Sigma-Aldrich, USA) as the manufacturer indicated. The ECM gel was prepared from Engelbreth-Holm-Swarm (EHS) mouse sarcoma and was composed of laminin as the major component, collagen type IV, heparan sulfate proteoglycan, entactin and other minor components. Cells were seeded at concentrations of 1,000,000 cells per well containing DMEM with antibiotics and 10% KSR. The plates were incubated at 37°C in a humidified atmosphere with 5% CO2. In the germ cell-removed group, the SSCs were removed from the in vitro culture using the differential plating method described by He et al, in which following 3 hr of incubation, somatic cells attached to the plate but SSCs had remained in the suspension and consequently were eliminated from the plate 26. No intervention was performed in the control group. To further evaluate the effectiveness of the differential plating method for removal of SSCs, the gene expression of spermatogonial markers was assessed. In this context, Thy1 and Bcl6b were used as the markers of undifferentiated spermatogonia 27 and ckit was used as a marker for differentiated spermatogonia 28. The medium was replaced with a fresh one every three days in both groups.
Evaluation of the colonization of SSCs: The colonization of SSCs in the control and germ cell-removed groups was assessed using an inverted microscope (IX71, Olympus, Japan).
RNA isolation and quantitative real-time PCR (qRT-PCR): Following trypsinization of the cultured cells (n=3 cell populations from different calves), the isolated cells were subjected to total RNA extraction using Trizol reagent (Fermentas, Germany). The extracted RNA was treated with DNase (Fermentas, Germany) to avoid DNA contamination. The concentration of RNA was determined using UV spectrophotometry (Eppendorff, Germany). The cDNAs were synthesized from 500 ng of RNA by oligo (dT) primers using RevertAid™ First Strand cDNA Synthesis Kit (Fermentas, Germany). Primers for genes of interest are presented in table 1. PCRs were performed using Master Mix and SYBR Green I (Fermentas, Germany) in an Applied Biosystems, StepOne™ thermal cycler (Applied Biosystems, USA). The PCR program started with an initial melting cycle for 5 min at 95°C to activate the polymerase, followed by 40 cycles of melting (30 s at 95°C), annealing (30 s at 58°C) and extension (30 s at 72°C). The quality of the PCR reactions was confirmed by melting curve analyses. For each sample, the reference gene (β-actin) and target gene were amplified in the same run. The target genes were normalized to the reference gene. The mean target gene threshold cycle (Ct) and mean exogenous control (β-actin) Ct for each sample were calculated from duplicate wells. The target gene threshold cycle (Ct) of the control was subtracted from the Ct of target gene, resulting in ∆Ct. In each experiment, the Ct of time-point 0 sample was considered as the calibrator. Subsequently, the ∆Ct of sample was then subtracted from the ∆Ct of calibrator, resulting in ∆∆Ct, which was used for calculation of the relative values of target gene expression for each sample 29.
Statistical analysis: Data were analyzed using MIXED procedure. In addition, LSMEANS statement was used to perform multiple comparisons. All analyses were conducted in SAS 30. Data are presented as mean ± SD. Differences with p<0.05 were considered significant.
Results :
Colonization of SCCs: While SSCs colonies were developed in the control group (Figure 1A), they were rarely observed in the germ cell-removed group (Figure 1B).
Gene expression: In the control group, the expression of Thy1 did not differ among days of culture (p>0.05; Figure 2). In the germ cell-removed group, Thy1 expression was not observed on days 6 and 12.
In the control group, the gene expression of Bcl6b was greater on days 6 (5.82±2.77 fold) and 12 (6.11±2.84 fold) compared with day 0 (p<0.05); however, it was not different between days 6 and 12 (p>0.05; Figure 2). The expression of Bcl6b was not observed in the germ cell-removed group on days 6 and 12.
The gene expression of ckit in the control group decreased 65% and 58% on days 6 and 12 of culture, respectively, as compared with day 0 (p<0.05), but it did not differ between days 6 and 12 (p>0.05; Figure 2). The expression of ckit was not observed in the germ cell-removed group.
In the control group, the expression of Gdnf did not change over the course of culture (p>0.05). In the germ cell-removed group, it was greater on days 6 (3.51±0.72) and 12 (4.14±0.65) than day 0 (p<0.01), but it was not different between days 6 and 12 (p>0.05). The gene expression of Gdnf did not differ between two groups on day 6; however, it was higher in the germ cell-removed than control group on day 12 (p<0.01; Figure 3).
The expression of Fgf2 was not different among days of culture in the control group (p>0.05). In the germ cell-removed group, it was higher on day 6 (3.72±1.09 fold) than day 0 (p=0.008), but it did not differ between days 0 and 12, and between days 6 and 12 (p>0.05). On day 6, the expression of Fgf2 in the germ cell-removed group was greater than that in the control group (p=0.031), but it did not differ between two groups on day 12 (p>0.05; Figure 4).
In the control group, the expression of Kitlg was higher on days 6 (8.11±3.14 fold) and 12 (5.61±1.53 fold) as compared with day 0 (p<0.05). In the germ cell-removed group, however, it did not differ among days of culture (p>0.05). The gene expression of Kitlg was greater in the control than germ cell-removed group on days 6 and 12 (p<0.01; Figure 5).
Discussion :
The present study was conducted to address whether SSCs removal could influence the molecular behavior of niche cells in terms of the expression of the main factors regulating SCCs development. SSCs colonies, resulting from the interactions between SSCs and Sertoli cells during in vitro culture 8,9, were rarely developed in the germ cell-removed group. Likewise, gene expression of Thy1, Bcl6b and ckit, as markers of spermatogonia 27,28, was not observed in the germ cell-removed group. Both of these findings indicate that SSC removal was successfully performed in the present study. Hence, the present study interestingly revealed that the expression of Gdnf increased during the culture in response to SSCs removal. Similarly, overexpression of GDNF and its mRNA has been reported following busulfan treatment, the drug which eliminates germ cells from seminiferous tubules in mice 19-21. GDNF overexpression has been reported to stimulate self-renewal and block differentiation in murine SSCs 2,31. On the other hand, mice lacking GDNF expression lose their undifferentiated spermatogonia with age, resulting in the presence of Sertoli cell only seminiferous tubules 2. Johnston et al has also shown the essential role of GDNF in SSCs replication and differentiation by assessing the gene expression of Gdnf over different stages of spermatogenesis in rat 6. The proliferative effect of GDNF on SSCs self-renewal has been substantiated by in vitro studies as well 5,9.
In addition, SSCs removal led to elevation in the gene expression of Fgf2, while the expression of Fgf2 was not affected over the course of culture in the control group. Similar to GDNF, contribution of FGF2 to SSCs self-renewal has been reported by in vitro 7 as well as in vivo 3 studies. FGF2 has also been suggested to have synergistic relationship with GDNF through upregulation of ETS variant 5 (Etv5), which positively impacts the expression of receptor tyrosine kinase Ret, mediating GDNF signals 3.
Although the gene expression of Kitlg in the germ cell-removed group did not change over the course of culture, it was lower as compared with that in the control group, implicating that the removal of SSCs abrogated the elevation in expression of Kitlg in response to exposure to ECM 17. Slight decrease in Kitlg mRNA expression has also been observed in mice treated with busulfan, albeit this reduction was not statistically significant 22. In contrast with GDNF and FGF2, KITLG is believed to be involved in SSCs differentiation. It has been indicated that Steel locus encodes soluble and membrane-bound forms of KITLG in mice 32. The membrane-bound domain of the Steel gene is deleted in Steel-dickie mutation 33, culminating in lack of KITLG production in Steel/Steeldickie mice 34. Steel/Steeldickie mice experience disrupted spermatogenesis and have testes with only primitive spermatogonia 4,34. Transduction of Sertoli cells with adenoviral and lentiviral vectors carrying the mouse Steel gene has been found to restore spermatogenesis 30,35. Furthermore, addition of recombinant KITLG to the tissue culture of testes obtained from Steel/Steeldickie mice would result in SSCs differentiation 4.
Taken together, it seems that removal of SSCs in the present study resulted in alterations in niche factors expression toward stimulation of self-renewal and abrogation of differentiation, by elevation in gene expression of Gdnf and Fgf2 as well as reduction in gene expression of Kitlg. Removal of SSCs from in vitro culture using differential plating partly resembles elimination of SSCs from seminiferous tubules by busulfan treatment 18. During the regeneration of germ cell population following testicular damage caused by busulfan, self-renewal is favored over differentiation 18. Hence, it could be surmised that somatic cells composing SSCs niche respond to SSCs elimination by upregulation of GDNF and FGF2 and downregulation of KITLG in order to repopulate niche by germ cells. Testicular dysfunction and germ cell depletion is common in male patients subjected to chemotherapy, which led to research to develop strategies to preserve the patient fertility 23. One of these strategies has been hormonal therapy either for ameliorating the testicular damage during chemotherapy or for enhancing the recovery after treatment 23. Data from the present study along with the finding of previous research 18-22 seem promising for potential application of the stimulatory or inhibitory effects of these growth factors to enhance the recovery of fertility in males with germ cell depletion due to testicular damage. Nevertheless, gene expression data in the present study requires to be initially confirmed using protein assays and then be trialed as treatment for recovery of fertility in males with testicular germ cell depletion.
In addition, these findings could be applied for enrichment of SSCs during in vitro culture prior to SSC transplantation in order to augment its efficiency 24. Previously, the beneficial effects of GDNF and FGF2 have been reported for enhancement of SSCs self-renewal 7,8. However, whether blockade of KITLG/cKIT pathway could accelerate the proliferatory effect of GDNF and FGF2 necessitates further studies to be addressed.
Considering the partly similar reaction of niche cells in response to either in vitro removal of SSCs using differential plating or in vivo elimination of SSCs using busulfan treatment 19-22, it could be suggested that removal of SSCs presented in the current research could serve as a model to study the events ensuing testicular germ cell damage and the mechanisms involved in regeneration of testicular germ cells afterwards.
Conclusion :
In conclusion, the present study showed that removal of SSCs would lead to increase in gene expression of GDNF and FGF2 and decrease in gene expression of KITLG, implicating that absence or decrease in the number of SSCs alter the behavior of niche cells toward stimulation of self-renewal and abrogation of differentiation, probably in order to restore germ cell population. These finding could help develop therapies for acceleration of male fertility recovery after chemotherapy and improve in vitro condition for enhancement of self-renewal prior to SSCs transplantation. Moreover, the results imply that SSCs removal from in vitro culture might serve as a model to study changes in niche following damage to germ cell population of testicular tissue.
Acknowledgement :
The present study was supported by Theriogenology Association, Faculty of Veterinary Medicine, University of Tehran. The authors would like to thank Mr. S. Pour Beiranvand for his kind assistance in the present study.
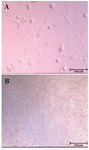

Figure 1. Colonization of SSCs in the control group; A) versus absence of colony formation in the germ-cell removed group; B) on day 12 of culture.
|
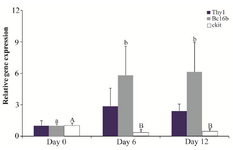
Figure 2. Relative gene expression of Thy1, Bcl6b and ckit in the control group (n=3) on days 0, 6 and 12. Different letters (a, b, A and B) indicate significant difference in the expression of the specified gene among timepoints (p<0.05).
|
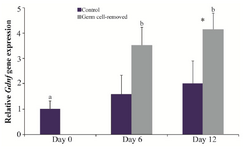
Figure 3. Relative gene expression of Gdnf in the control and germ cell-removed groups (n=3) on days 0, 6 and 12. Different letters (a and b) indicate significant difference within groups between different timepoints (p<0.05). Asterisk (*) indicates significant difference between two experimental groups at the specified timepoint (p<0.05).
|
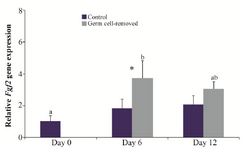
Figure 4. Relative gene expression of Fgf2 in the control and germ cell-removed groups (n=3) on days 0, 6 and 12. Different letters (a and b) indicate significant difference within groups between different timepoints (p<0.05). Asterisk (*) indicates significant difference between two experimental groups at the specified timepoint (p<0.05).
|
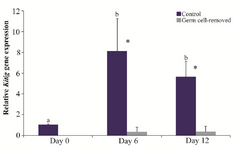
Figure 5. Relative gene expression of Kitlg in the control and germ cell-removed groups (n=3) on days 0, 6 and 12. Different letters (a and b) indicate significant difference within groups between different timepoints (p<0.05). Asterisks (*) indicate significant difference between two experimental groups at the specified timepoint (p<0.05).
|
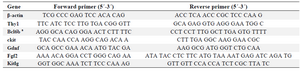
Table 1. Primer sequences used for qRT-PCR.
a: B-cell CLL/lymphoma 6, member B.
|
|